সংক্ৰমণ
| এইটো এটা অনূদিত বা অনুবাদ চলি থকা প্ৰবন্ধ। ইয়াৰ তাৎপৰ্য্য কেইবাটাও হ'ব পাৰে। অধিক জানিবৰ বাবে সোঁফালে থকা [দেখুৱাওক] লিংকটোত ক্লিক কৰক।
ওপৰত দিয়াৰ দৰে বিভিন্ন কাৰণত অনুবাদ প্ৰবন্ধ এটা দেখাত আপচু বা চুটি দেখা যাব পাৰে। লগতে প্ৰবন্ধত সম্পূৰ্ণ তথ্য নাথাকিবও পাৰে। আধৰুৱা তথ্য ক্ষতিকাৰক হ'ব পাৰে। সেয়ে এনে প্ৰবন্ধ পঢ়োঁতে পঢ়ুৱৈক নিজৰ বিবেক ব্যৱহাৰ কৰি প্ৰয়োজন হ'লে আমাৰ ৱিকিপিডিয়াতে ক্ষান্ত নাথাকি মূল প্ৰবন্ধটোও বিচাৰি পঢ়ি ল'বলৈ অনুৰোধ কৰা হ'ল। লগতে অনুবাদকসকলকো সাবধান হ'বলৈ অনুৰোধ কৰা হ'ল। |
| সংক্ৰামক ৰোগ | |
|---|---|
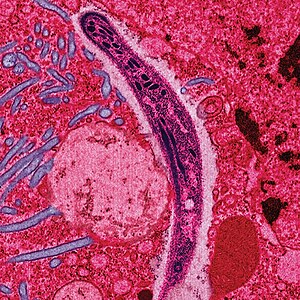 | |
| ৰং কৰা এক ইলেক্ট্ৰন মাইক্ৰ'গ্ৰাফত এটা মেলেৰিয়া স্প'ৰ'জইট পেটৰ ভিতৰৰ আৱৰণীৰ মাজেৰে পাৰ হোৱা দেখা গৈছে। | |
| বিভাগ | সংক্ৰামক ৰোগ |
সংক্ৰমণ বুলিলে জীৱৰ দেহত ৰোগ সৃষ্টিকাৰী বীজাণুৱে কৰা আক্ৰমণ, সংখ্যা বৃদ্ধি আৰু বীজাণু বা ইয়াৰ বিষাক্ত পদাৰ্থৰ প্ৰতি দেহৰ কলাৰ সঁহাৰিক বুজায়।[1][2] সংক্ৰমণৰ পৰা হোৱা ৰোগক সংক্ৰামক ৰোগ বোলে।
ভাইৰাছ, ভিৰইড, প্ৰিয়ন, ৰোগ সৃষ্টিকাৰী বেক্টেৰিয়া, পৰজীৱী যেনে পেলু বা কৃমি, সন্ধিপদী প্ৰাণী যেনে চিকৰা, চাহী, ভেঁকুৰ যেনে খৰ আদিয়ে সংক্ৰমণ কৰে।
জীৱই নিজৰ প্ৰতিৰক্ষা প্ৰণালী ব্যৱহাৰ কৰি সংক্ৰমণ ৰোধ কৰিব পাৰে। স্তন্যপায়ী প্ৰাণীয়ে সহজাত প্ৰতিৰক্ষাৰে সংক্ৰমণ ৰোধ কৰিবলৈ চেষ্টা কৰে যাৰ ফলত প্ৰায়েই প্ৰদাহ হৈ পাছলৈ লব্ধ প্ৰতিৰক্ষা সক্ৰিয় হয়।[3]
সংক্ৰমণৰ চিকিৎসাৰ বাবে নিৰ্দিষ্ট ঔষধ যেনে এণ্টিবায়'টিক, এণ্টিভাইৰেল, এণ্টিফাংগেল, এণ্টিপ্ৰট'জ'ৱেল আৰু এণ্টিহেল্মিন্থিক্ছ ব্যৱহাৰ কৰা হয়। ২০১৩ চনত সংক্ৰমণৰ ফলত প্ৰায় ৯২ লাখ মানুহৰ মৃত্যু হৈছিল (সকলো মৃত্যুৰ প্ৰয় ১৭%)।[4] সংক্ৰমণক প্ৰাধান্য দিয়া চিকিৎসা বিজ্ঞানৰ শাখাটোক সংক্ৰামক ৰোগ বোলে।[5]
কম সময়ৰ বাবে হোৱা সংক্ৰমণক "একিউট" (acute) বা হস্ৰম্যাদী সংক্ৰমণ বোলে। বেছি দিনৰ বাবে হোৱা সংক্ৰমণক "ক্ৰ'নিক" (chronic) বা দীৰ্ঘম্যাদী সংক্ৰমণ বোলে। কাৰকৰ ওপৰত ভিত্তি কৰি (যেনে বেক্টেৰিয়া, ভাইৰাছ, ভেঁকুৰ, পৰজীৱী) আৰু প্ৰণালীগত লক্ষণ থকা-নথকাক লৈও সংক্ৰমণৰ শ্ৰেণীবিভাজন কৰিব পাৰি।
প্ৰাথমিক বনাম সুযোগ সন্ধানী
[সম্পাদনা কৰক]জীৱাণু অসংখ্য প্ৰকাৰ থাকিলেও সুস্থ মানুহৰ দেহত কেইটামানেহে ৰোগৰ সৃষ্টি কৰিব পাৰে।[6] জীৱাণুবিধে দেহত কৰা আক্ৰমণ আৰু প্ৰতিৰক্ষা প্ৰণালীৰ প্ৰত্যাক্ৰমণৰ ওপৰত সংক্ৰমণৰ প্ৰাবল্য নিৰ্ভৰ কৰে। প্ৰতিৰক্ষা প্ৰণালীয়েও সংক্ৰমণ ৰোধ কৰিবলৈ গৈ দেহৰ ক্ষতি কৰিব পাৰে। সেয়েহে প্ৰতিৰক্ষা প্ৰণালীৰ স্থিতিলৈ চাই জীৱাণুবোৰক প্ৰাথমিক জীৱাণু আৰু সুযোগ সন্ধানী জীৱাণু বুলি ভাগ কৰা হয়।
- প্ৰাথমিক জীৱাণু
- প্ৰাথমিক জীৱাণুবোৰে সুস্থ প্ৰাণীৰ দেহত ৰোগৰ সৃষ্টি কৰিব পাৰে। এনে ৰোগৰ তীব্ৰতা সেই বীজাণুবোৰৰ সংখ্যা-বৃদ্ধি আৰু ব্যাপ্তিৰ ওপৰত নিৰ্ভৰ কৰে। মানুহৰ বহু প্ৰাথমিক বীজাণুৱে কেৱল মানুহকে সংক্ৰমিত কৰে। কিন্তু পৰিবেশ বা অন্য প্ৰাণীক সংক্ৰমণ কৰা বীজাণুৱেও মানুহৰ কেইবাটাও জটিল ৰোগ কৰিব পাৰে।
- সুযোগ সন্ধানী জীৱাণু
- সুযোগ সন্ধানী জীৱাণুৱে প্ৰতিৰোধ ক্ষমতা কম হ'লে বা অস্বাভাৱিক প্ৰক্ৰিয়াৰে শৰীৰত প্ৰৱেশ কৰিব পাৰিলে (যেনে আঘাত) ৰোগৰ সৃষ্টি কৰে। Opportunistic infection may be caused by microbes ordinarily in contact with the host, such as pathogenic bacteria or fungi in the gastrointestinal or the upper respiratory tract, and they may also result from (otherwise innocuous) microbes acquired from other hosts (as in Clostridium difficile colitis) or from the environment as a result of traumatic introduction (as in surgical wound infections or compound fractures). An opportunistic disease requires impairment of host defenses, which may occur as a result of genetic defects (such as Chronic granulomatous disease), exposure to antimicrobial drugs or immunosuppressive chemicals (as might occur following poisoning or cancer chemotherapy), exposure to ionizing radiation, or as a result of an infectious disease with immunosuppressive activity (such as with measles, malaria or HIV disease). Primary pathogens may also cause more severe disease in a host with depressed resistance than would normally occur in an immunosufficient host.[7]
- প্ৰাথমিক সংক্ৰমণ বনাম গৌণ সংক্ৰমণ
- প্ৰাথমিক সংক্ৰমণক স্বাস্থ্যৰ সমস্যাৰ মূল কাৰণ হিচাপে গণ্য কৰিব পাৰি। ইয়াৰ বিপৰীতে অপ্ৰধান সংক্ৰমণ মূল কাৰণৰ জটিলতাৰ পাছতহে হয়। যেনে- হাওঁফাওঁৰ যক্ষ্মাৰ প্ৰায়েই এটা প্ৰাথমিক সংক্ৰমণ, কিন্তু জুই লগাৰ পাছত বা দ আঘাত পোৱাৰ পাছত বীজাণু গাত প্ৰৱেশ কৰিলে সেয়া গৌণ সংক্ৰমণ। প্ৰাথমিক জীৱাণুবোৰে প্ৰায়েই প্ৰাথমিক সংক্ৰমণ আৰু গৌণ সংক্ৰমণ দুয়োটাই কৰে। সাধাৰণতে সুযোগ সন্ধানী সংক্ৰমণক গৌণ সংক্ৰমণ বুলি গণ্য কৰা হয়।
সংক্ৰমণ হয় নে নহয়
[সম্পাদনা কৰক]One way of proving that a given disease is "infectious", is to satisfy Koch's postulates (first proposed by Robert Koch), which demands that the infectious agent be identified only in patients and not in healthy controls, and that patients who contract the agent also develop the disease. These postulates were first used in the discovery that Mycobacteria species cause tuberculosis. Koch's postulates can not be applied ethically for many human diseases because they require experimental infection of a healthy individual with a pathogen produced as a pure culture. Often, even clearly infectious diseases do not meet the infectious criteria. For example, Treponema pallidum, the causative spirochete of syphilis, cannot be cultured in vitro – however the organism can be cultured in rabbit testes. It is less clear that a pure culture comes from an animal source serving as host than it is when derived from microbes derived from plate culture. Epidemiology is another important tool used to study disease in a population. For infectious diseases it helps to determine if a disease outbreak is sporadic (occasional occurrence), endemic (regular cases often occurring in a region), epidemic (an unusually high number of cases in a region), or pandemic (a global epidemic).
সোঁচৰা
[সম্পাদনা কৰক]Infectious diseases are sometimes called contagious disease when they are easily transmitted by contact with an ill person or their secretions (e.g., influenza). Thus, a contagious disease is a subset of infectious disease that is especially infective or easily transmitted. Other types of infectious/transmissible/communicable diseases with more specialized routes of infection, such as vector transmission or sexual transmission, are usually not regarded as "contagious", and often do not require medical isolation (sometimes loosely called quarantine) of victims. However, this specialized connotation of the word "contagious" and "contagious disease" (easy transmissibility) is not always respected in popular use. Infectious diseases are commonly transmitted from person to person through direct contact. The types of contact are through person to person and droplet spread. Indirect contact such as airborne transmission, contaminated objects, food and drinking water, animal person contact, animal reservoirs, insect bites, and environmental reservoirs are another way infectious diseases are transmitted,[8]
শৰীৰৰ স্থান অনুসৰি
[সম্পাদনা কৰক]Infections can be classified by the anatomic location or organ system infected, including:
- Urinary tract infection
- Skin infection
- Respiratory tract infection
- Odontogenic infection (an infection that originates within a tooth or in the closely surrounding tissues)
- Vaginal infections
- Intra-amniotic infection
In addition, locations of inflammation where infection is the most common cause include pneumonia, meningitis and salpingitis.
লক্ষণ আৰু চিহ্ন
[সম্পাদনা কৰক]The symptoms of an infection depend on the type of disease. Some signs of infection affect the whole body generally, such as fatigue, loss of appetite, weight loss, fevers, night sweats, chills, aches and pains. Others are specific to individual body parts, such as skin rashes, coughing, or a runny nose.
In certain cases, infectious diseases may be asymptomatic for much or even all of their course in a given host. In the latter case, the disease may only be defined as a "disease" (which by definition means an illness) in hosts who secondarily become ill after contact with an asymptomatic carrier. An infection is not synonymous with an infectious disease, as some infections do not cause illness in a host.[7]
বেক্টেৰিয়াজনিত বা ভাইৰাছজনিত
[সম্পাদনা কৰক]Bacterial and viral infections can both cause the same kinds of symptoms. It can be difficult to distinguish which is the cause of a specific infection.[9] It's important to distinguish, because viral infections cannot be cured by antibiotics.[10]
| Characteristic | Viral infection | Bacterial infection |
|---|---|---|
| Typical symptoms | In general, viral infections are systemic. This means they involve many different parts of the body or more than one body system at the same time; i.e. a runny nose, sinus congestion, cough, body aches etc. They can be local at times as in viral conjunctivitis or "pink eye" and herpes. Only a few viral infections are painful, like herpes. The pain of viral infections is often described as itchy or burning.[9] | The classic symptoms of a bacterial infection are localized redness, heat, swelling and pain. One of the hallmarks of a bacterial infection is local pain, pain that is in a specific part of the body. For example, if a cut occurs and is infected with bacteria, pain occurs at the site of the infection. Bacterial throat pain is often characterized by more pain on one side of the throat. An ear infection is more likely to be diagnosed as bacterial if the pain occurs in only one ear.[9] A cut that produces pus and milky-colored liquid is most likely [স্পষ্টীকৰণৰ প্ৰয়োজন][11] |
| Cause | Pathogenic viruses | Pathogenic bacteria |
ৰোগ প্ৰক্ৰিয়া
[সম্পাদনা কৰক]There is a general chain of events that applies to infections.[12] The chain of events involves several steps—which include the infectious agent, reservoir, entering a susceptible host, exit and transmission to new hosts. Each of the links must be present in a chronological order for an infection to develop. Understanding these steps helps health care workers target the infection and prevent it from occurring in the first place.[13]
Colonization
[সম্পাদনা কৰক]
Infection begins when an organism successfully enters the body, grows and multiplies. This is referred to as colonization. Most humans are not easily infected. Those who are weak, sick, malnourished, have cancer or are diabetic have increased susceptibility to chronic or persistent infections. Individuals who have a suppressed immune system are particularly susceptible to opportunistic infections. Entrance to the host at host-pathogen interface, generally occurs through the mucosa in orifices like the oral cavity, nose, eyes, genitalia, anus, or the microbe can enter through open wounds. While a few organisms can grow at the initial site of entry, many migrate and cause systemic infection in different organs. Some pathogens grow within the host cells (intracellular) whereas others grow freely in bodily fluids.
Wound colonization refers to nonreplicating microorganisms within the wound, while in infected wounds, replicating organisms exist and tissue is injured. All multicellular organisms are colonized to some degree by extrinsic organisms, and the vast majority of these exist in either a mutualistic or commensal relationship with the host. An example of the former is the anaerobic bacteria species, which colonizes the mammalian colon, and an example of the latter is various species of staphylococcus that exist on human skin. Neither of these colonizations are considered infections. The difference between an infection and a colonization is often only a matter of circumstance. Non-pathogenic organisms can become pathogenic given specific conditions, and even the most virulent organism requires certain circumstances to cause a compromising infection. Some colonizing bacteria, such as Corynebacteria sp. and viridans streptococci, prevent the adhesion and colonization of pathogenic bacteria and thus have a symbiotic relationship with the host, preventing infection and speeding wound healing.

The variables involved in the outcome of a host becoming inoculated by a pathogen and the ultimate outcome include:
- the route of entry of the pathogen and the access to host regions that it gains
- the intrinsic virulence of the particular organism
- the quantity or load of the initial inoculant
- the immune status of the host being colonized
As an example, several staphylococcal species remain harmless on the skin, but, when present in a normally sterile space, such as in the capsule of a joint or the peritoneum, multiply without resistance and cause harm.
An interesting fact that gas chromatography–mass spectrometry, 16S ribosomal RNA analysis, omics, and other advanced technologies have made more apparent to humans in recent decades is that microbial colonization is very common even in environments that humans think of as being nearly sterile. Because it is normal to have bacterial colonization, it is difficult to know which chronic wounds can be classified as infected and how much risk of progression exists. Despite the huge number of wounds seen in clinical practice, there are limited quality data for evaluated symptoms and signs. A review of chronic wounds in the Journal of the American Medical Association's "Rational Clinical Examination Series" quantified the importance of increased pain as an indicator of infection.[17] The review showed that the most useful finding is an increase in the level of pain [likelihood ratio (LR) range, 11–20] makes infection much more likely, but the absence of pain (negative likelihood ratio range, 0.64–0.88) does not rule out infection (summary LR 0.64–0.88).
Disease
[সম্পাদনা কৰক]Disease can arise if the host's protective immune mechanisms are compromised and the organism inflicts damage on the host. Microorganisms can cause tissue damage by releasing a variety of toxins or destructive enzymes. For example, Clostridium tetani releases a toxin that paralyzes muscles, and staphylococcus releases toxins that produce shock and sepsis. Not all infectious agents cause disease in all hosts. For example, less than 5% of individuals infected with polio develop disease.[18] On the other hand, some infectious agents are highly virulent. The prion causing mad cow disease and Creutzfeldt–Jakob disease invariably kills all animals and people that are infected.
Persistent infections occur because the body is unable to clear the organism after the initial infection. Persistent infections are characterized by the continual presence of the infectious organism, often as latent infection with occasional recurrent relapses of active infection. There are some viruses that can maintain a persistent infection by infecting different cells of the body. Some viruses once acquired never leave the body. A typical example is the herpes virus, which tends to hide in nerves and become reactivated when specific circumstances arise.
Persistent infections cause millions of deaths globally each year.[19] Chronic infections by parasites account for a high morbidity and mortality in many underdeveloped countries.
Transmission
[সম্পাদনা কৰক]For infecting organisms to survive and repeat the infection cycle in other hosts, they (or their progeny) must leave an existing reservoir and cause infection elsewhere. Infection transmission can take place via many potential routes:
- Droplet contact, also known as the respiratory route, and the resultant infection can be termed airborne disease. If an infected person coughs or sneezes on another person the microorganisms, suspended in warm, moist droplets, may enter the body through the nose, mouth or eye surfaces.
- Fecal-oral transmission, wherein foodstuffs or water become contaminated (by people not washing their hands before preparing food, or untreated sewage being released into a drinking water supply) and the people who eat and drink them become infected. Common fecal-oral transmitted pathogens include Vibrio cholerae, Giardia species, rotaviruses, Entameba histolytica, Escherichia coli, and tape worms.[20] Most of these pathogens cause gastroenteritis.
- Sexual transmission, with the resulting disease being called sexually transmitted disease
- Oral transmission, Diseases that are transmitted primarily by oral means may be caught through direct oral contact such as kissing, or by indirect contact such as by sharing a drinking glass or a cigarette.
- Transmission by direct contact, Some diseases that are transmissible by direct contact include athlete's foot, impetigo and warts
- Vehicle Transmission, transmission by an inanimate reservoir (food, water, soil).[21]
- Vertical transmission, directly from the mother to an embryo, fetus or baby during pregnancy or childbirth. It can occur when the mother gets an infection as an intercurrent disease in pregnancy.
- Iatrogenic transmission, due to medical procedures such as injection or transplantation of infected material.

- Vector-borne transmission, transmitted by a vector, which is an organism that does not cause disease itself but that transmits infection by conveying pathogens from one host to another.[22]
The relationship between virulence versus transmissibility is complex; if a disease is rapidly fatal, the host may die before the microbe can be passed along to another host.
ৰোগ নিৰ্ণয়
[সম্পাদনা কৰক]Diagnosis of infectious disease sometimes involves identifying an infectious agent either directly or indirectly. In practice most minor infectious diseases such as warts, cutaneous abscesses, respiratory system infections and diarrheal diseases are diagnosed by their clinical presentation and treated without knowledge of the specific causative agent. Conclusions about the cause of the disease are based upon the likelihood that a patient came in contact with a particular agent, the presence of a microbe in a community, and other epidemiological considerations. Given sufficient effort, all known infectious agents can be specifically identified. The benefits of identification, however, are often greatly outweighed by the cost, as often there is no specific treatment, the cause is obvious, or the outcome of an infection is benign.
Diagnosis of infectious disease is nearly always initiated by medical history and physical examination. More detailed identification techniques involve the culture of infectious agents isolated from a patient. Culture allows identification of infectious organisms by examining their microscopic features, by detecting the presence of substances produced by pathogens, and by directly identifying an organism by its genotype. Other techniques (such as X-rays, CAT scans, PET scans or NMR) are used to produce images of internal abnormalities resulting from the growth of an infectious agent. The images are useful in detection of, for example, a bone abscess or a spongiform encephalopathy produced by a prion.
Symptomatic diagnostics
[সম্পাদনা কৰক]The diagnosis is aided by the presenting symptoms in any individual with an infectious disease, yet it usually needs additional diagnostic techniques to confirm the suspicion. Some signs are specifically characteristic and indicative of a disease and are called pathognomonic signs; but these are rare. Not all infections are symptomatic.[23]
In children the presence of cyanosis, rapid breathing, poor peripheral perfusion, or a petechial rash increases the risk of a serious infection by greater than 5 fold.[24] Other important indicators include parental concern, clinical instinct, and temperature greater than 40 °C.[24]
Microbial culture
[সম্পাদনা কৰক]
Microbiological culture is a principal tool used to diagnose infectious disease. In a microbial culture, a growth medium is provided for a specific agent. A sample taken from potentially diseased tissue or fluid is then tested for the presence of an infectious agent able to grow within that medium. Most pathogenic bacteria are easily grown on nutrient agar, a form of solid medium that supplies carbohydrates and proteins necessary for growth of a bacterium, along with copious amounts of water. A single bacterium will grow into a visible mound on the surface of the plate called a colony, which may be separated from other colonies or melded together into a "lawn". The size, color, shape and form of a colony is characteristic of the bacterial species, its specific genetic makeup (its strain), and the environment that supports its growth. Other ingredients are often added to the plate to aid in identification. Plates may contain substances that permit the growth of some bacteria and not others, or that change color in response to certain bacteria and not others. Bacteriological plates such as these are commonly used in the clinical identification of infectious bacterium. Microbial culture may also be used in the identification of viruses: the medium in this case being cells grown in culture that the virus can infect, and then alter or kill. In the case of viral identification, a region of dead cells results from viral growth, and is called a "plaque". Eukaryotic parasites may also be grown in culture as a means of identifying a particular agent.
In the absence of suitable plate culture techniques, some microbes require culture within live animals. Bacteria such as Mycobacterium leprae and Treponema pallidum can be grown in animals, although serological and microscopic techniques make the use of live animals unnecessary. Viruses are also usually identified using alternatives to growth in culture or animals. Some viruses may be grown in embryonated eggs. Another useful identification method is Xenodiagnosis, or the use of a vector to support the growth of an infectious agent. Chagas disease is the most significant example, because it is difficult to directly demonstrate the presence of the causative agent, Trypanosoma cruzi in a patient, which therefore makes it difficult to definitively make a diagnosis. In this case, xenodiagnosis involves the use of the vector of the Chagas agent T. cruzi, an uninfected triatomine bug, which takes a blood meal from a person suspected of having been infected. The bug is later inspected for growth of T. cruzi within its gut.
Microscopy
[সম্পাদনা কৰক]Another principal tool in the diagnosis of infectious disease is microscopy. Virtually all of the culture techniques discussed above rely, at some point, on microscopic examination for definitive identification of the infectious agent. Microscopy may be carried out with simple instruments, such as the compound light microscope, or with instruments as complex as an electron microscope. Samples obtained from patients may be viewed directly under the light microscope, and can often rapidly lead to identification. Microscopy is often also used in conjunction with biochemical staining techniques, and can be made exquisitely specific when used in combination with antibody based techniques. For example, the use of antibodies made artificially fluorescent (fluorescently labeled antibodies) can be directed to bind to and identify a specific antigens present on a pathogen. A fluorescence microscope is then used to detect fluorescently labeled antibodies bound to internalized antigens within clinical samples or cultured cells. This technique is especially useful in the diagnosis of viral diseases, where the light microscope is incapable of identifying a virus directly.
Other microscopic procedures may also aid in identifying infectious agents. Almost all cells readily stain with a number of basic dyes due to the electrostatic attraction between negatively charged cellular molecules and the positive charge on the dye. A cell is normally transparent under a microscope, and using a stain increases the contrast of a cell with its background. Staining a cell with a dye such as Giemsa stain or crystal violet allows a microscopist to describe its size, shape, internal and external components and its associations with other cells. The response of bacteria to different staining procedures is used in the taxonomic classification of microbes as well. Two methods, the Gram stain and the acid-fast stain, are the standard approaches used to classify bacteria and to diagnosis of disease. The Gram stain identifies the bacterial groups Firmicutes and Actinobacteria, both of which contain many significant human pathogens. The acid-fast staining procedure identifies the Actinobacterial genera Mycobacterium and Nocardia.
Biochemical tests
[সম্পাদনা কৰক]Biochemical tests used in the identification of infectious agents include the detection of metabolic or enzymatic products characteristic of a particular infectious agent. Since bacteria ferment carbohydrates in patterns characteristic of their genus and species, the detection of fermentation products is commonly used in bacterial identification. Acids, alcohols and gases are usually detected in these tests when bacteria are grown in selective liquid or solid media.
The isolation of enzymes from infected tissue can also provide the basis of a biochemical diagnosis of an infectious disease. For example, humans can make neither RNA replicases nor reverse transcriptase, and the presence of these enzymes are characteristic of specific types of viral infections. The ability of the viral protein hemagglutinin to bind red blood cells together into a detectable matrix may also be characterized as a biochemical test for viral infection, although strictly speaking hemagglutinin is not an enzyme and has no metabolic function.
Serological methods are highly sensitive, specific and often extremely rapid tests used to identify microorganisms. These tests are based upon the ability of an antibody to bind specifically to an antigen. The antigen, usually a protein or carbohydrate made by an infectious agent, is bound by the antibody. This binding then sets off a chain of events that can be visibly obvious in various ways, dependent upon the test. For example, "Strep throat" is often diagnosed within minutes, and is based on the appearance of antigens made by the causative agent, S. pyogenes, that is retrieved from a patients throat with a cotton swab. Serological tests, if available, are usually the preferred route of identification, however the tests are costly to develop and the reagents used in the test often require refrigeration. Some serological methods are extremely costly, although when commonly used, such as with the "strep test", they can be inexpensive.[7]
Complex serological techniques have been developed into what are known as Immunoassays. Immunoassays can use the basic antibody – antigen binding as the basis to produce an electro-magnetic or particle radiation signal, which can be detected by some form of instrumentation. Signal of unknowns can be compared to that of standards allowing quantitation of the target antigen. To aid in the diagnosis of infectious diseases, immunoassays can detect or measure antigens from either infectious agents or proteins generated by an infected organism in response to a foreign agent. For example, immunoassay A may detect the presence of a surface protein from a virus particle. Immunoassay B on the other hand may detect or measure antibodies produced by an organism's immune system that are made to neutralize and allow the destruction of the virus.
Instrumentation can be used to read extremely small signals created by secondary reactions linked to the antibody – antigen binding. Instrumentation can control sampling, reagent use, reaction times, signal detection, calculation of results, and data management to yield a cost effective automated process for diagnosis of infectious disease.
PCR-based diagnostics
[সম্পাদনা কৰক]Technologies based upon the polymerase chain reaction (PCR) method will become nearly ubiquitous gold standards of diagnostics of the near future, for several reasons. First, the catalog of infectious agents has grown to the point that virtually all of the significant infectious agents of the human population have been identified. Second, an infectious agent must grow within the human body to cause disease; essentially it must amplify its own nucleic acids in order to cause a disease. This amplification of nucleic acid in infected tissue offers an opportunity to detect the infectious agent by using PCR. Third, the essential tools for directing PCR, primers, are derived from the genomes of infectious agents, and with time those genomes will be known, if they are not already.
Thus, the technological ability to detect any infectious agent rapidly and specifically are currently available. The only remaining blockades to the use of PCR as a standard tool of diagnosis are in its cost and application, neither of which is insurmountable. The diagnosis of a few diseases will not benefit from the development of PCR methods, such as some of the clostridial diseases (tetanus and botulism). These diseases are fundamentally biological poisonings by relatively small numbers of infectious bacteria that produce extremely potent neurotoxins. A significant proliferation of the infectious agent does not occur, this limits the ability of PCR to detect the presence of any bacteria.
Metagenomic sequencing
[সম্পাদনা কৰক]Given the wide range of bacteria, viruses, and other pathogens that cause debilitating and life-threatening illness, the ability to quickly identify the cause of infection is important yet often challenging. For example, more than half of cases of encephalitis, a severe illness affecting the brain, remain undiagnosed, despite extensive testing using state-of-the-art clinical laboratory methods. Metagenomics is currently being researched for clinical use, and shows promise as a sensitive and rapid way to diagnose infection using a single all-encompassing test. This test is similar to current PCR tests; however, amplification of genetic material is unbiased rather than using primers for a specific infectious agent. This amplification step is followed by next-generation sequencing and alignment comparisons using large databases of thousands of organismic and viral genomes.
Metagenomic sequencing could prove especially useful for diagnosis when the patient is immunocompromised. An ever-wider array of infectious agents can cause serious harm to individuals with immunosuppression, so clinical screening must often be broader. Additionally, the expression of symptoms is often atypical, making clinical diagnosis based on presentation more difficult. Thirdly, diagnostic methods that rely on the detection of antibodies are more likely to fail. A broad, sensitive test for pathogens that detects the presence of infectious material rather than antibodies is therefore highly desirable.
Indication of tests
[সম্পাদনা কৰক]There is usually an indication for a specific identification of an infectious agent only when such identification can aid in the treatment or prevention of the disease, or to advance knowledge of the course of an illness prior to the development of effective therapeutic or preventative measures. For example, in the early 1980s, prior to the appearance of AZT for the treatment of AIDS, the course of the disease was closely followed by monitoring the composition of patient blood samples, even though the outcome would not offer the patient any further treatment options. In part, these studies on the appearance of HIV in specific communities permitted the advancement of hypotheses as to the route of transmission of the virus. By understanding how the disease was transmitted, resources could be targeted to the communities at greatest risk in campaigns aimed at reducing the number of new infections. The specific serological diagnostic identification, and later genotypic or molecular identification, of HIV also enabled the development of hypotheses as to the temporal and geographical origins of the virus, as well as a myriad of other hypothesis.[7] The development of molecular diagnostic tools have enabled physicians and researchers to monitor the efficacy of treatment with anti-retroviral drugs. Molecular diagnostics are now commonly used to identify HIV in healthy people long before the onset of illness and have been used to demonstrate the existence of people who are genetically resistant to HIV infection. Thus, while there still is no cure for AIDS, there is great therapeutic and predictive benefit to identifying the virus and monitoring the virus levels within the blood of infected individuals, both for the patient and for the community at large.
নিবাৰণ
[সম্পাদনা কৰক]
Techniques like hand washing, wearing gowns, and wearing face masks can help prevent infections from being passed from one person to another. Aseptic technique was introduced in medicine and surgery in the late 19th century and greatly reduced the incidence of infections caused by surgery. Frequent hand washing remains the most important defense against the spread of unwanted organisms.[26] There are other forms of prevention such as avoiding the use of illicit drugs, using a condom, and having a healthy lifestyle with a balanced diet and regular exercise. Cooking foods well and avoiding foods that have been left outside for a long time is also important.
Antimicrobial substances used to prevent transmission of infections include:
- antiseptics, which are applied to living tissue/skin
- disinfectants, which destroy microorganisms found on non-living objects.
- antibiotics, called prophylactic when given as prevention rather as treatment of infection. However, long term use of antibiotics leads to resistance of bacteria. While humans do not become immune to antibiotics, the bacteria does. Thus, avoiding using antibiotics longer than necessary helps preventing bacteria from forming mutations that aide in antibiotic resistance.
One of the ways to prevent or slow down the transmission of infectious diseases is to recognize the different characteristics of various diseases.[27] Some critical disease characteristics that should be evaluated include virulence, distance traveled by victims, and level of contagiousness. The human strains of Ebola virus, for example, incapacitate their victims extremely quickly and kill them soon after. As a result, the victims of this disease do not have the opportunity to travel very far from the initial infection zone.[28] Also, this virus must spread through skin lesions or permeable membranes such as the eye. Thus, the initial stage of Ebola is not very contagious since its victims experience only internal hemorrhaging. As a result of the above features, the spread of Ebola is very rapid and usually stays within a relatively confined geographical area. In contrast, the Human Immunodeficiency Virus (HIV) kills its victims very slowly by attacking their immune system.[7] As a result, many of its victims transmit the virus to other individuals before even realizing that they are carrying the disease. Also, the relatively low virulence allows its victims to travel long distances, increasing the likelihood of an epidemic.
Another effective way to decrease the transmission rate of infectious diseases is to recognize the effects of small-world networks.[27] In epidemics, there are often extensive interactions within hubs or groups of infected individuals and other interactions within discrete hubs of susceptible individuals. Despite the low interaction between discrete hubs, the disease can jump to and spread in a susceptible hub via a single or few interactions with an infected hub. Thus, infection rates in small-world networks can be reduced somewhat if interactions between individuals within infected hubs are eliminated (Figure 1). However, infection rates can be drastically reduced if the main focus is on the prevention of transmission jumps between hubs. The use of needle exchange programs in areas with a high density of drug users with HIV is an example of the successful implementation of this treatment method. [6] Another example is the use of ring culling or vaccination of potentially susceptible livestock in adjacent farms to prevent the spread of the foot-and-mouth virus in 2001.[29]
A general method to prevent transmission of vector-borne pathogens is pest control.
Immunity
[সম্পাদনা কৰক]
Infection with most pathogens does not result in death of the host and the offending organism is ultimately cleared after the symptoms of the disease have waned.[6] This process requires immune mechanisms to kill or inactivate the inoculum of the pathogen. Specific acquired immunity against infectious diseases may be mediated by antibodies and/or T lymphocytes. Immunity mediated by these two factors may be manifested by:
- a direct effect upon a pathogen, such as antibody-initiated complement-dependent bacteriolysis, opsonoization, phagocytosis and killing, as occurs for some bacteria,
- neutralization of viruses so that these organisms cannot enter cells,
- or by T lymphocytes, which will kill a cell parasitized by a microorganism.
The immune system response to a microorganism often causes symptoms such as a high fever and inflammation, and has the potential to be more devastating than direct damage caused by a microbe.[7]
Resistance to infection (immunity) may be acquired following a disease, by asymptomatic carriage of the pathogen, by harboring an organism with a similar structure (crossreacting), or by vaccination. Knowledge of the protective antigens and specific acquired host immune factors is more complete for primary pathogens than for opportunistic pathogens. There is also the phenomenon of herd immunity which offers a measure of protection to those otherwise vulnerable people when a large enough proportion of the population has acquired immunity from certain infections.
Immune resistance to an infectious disease requires a critical level of either antigen-specific antibodies and/or T cells when the host encounters the pathogen. Some individuals develop natural serum antibodies to the surface polysaccharides of some agents although they have had little or no contact with the agent, these natural antibodies confer specific protection to adults and are passively transmitted to newborns.
Host genetic factors
[সম্পাদনা কৰক]The organism that is the target of an infecting action of a specific infectious agent is called the host. The host harbors and agent in a mature, or sexually active stage phase called the definitive host. The intermediate host comes in contact during the larvae stage. A host can be anything living and can attain to asexual and sexual reproduction.[30] The clearance of the pathogens, either treatment-induced or spontaneous, it can be influenced by the genetic variants carried by the individual patients. For instance, for genotype 1 hepatitis C treated with Pegylated interferon-alpha-2a or Pegylated interferon-alpha-2b (brand names Pegasys or PEG-Intron) combined with ribavirin, it has been shown that genetic polymorphisms near the human IL28B gene, encoding interferon lambda 3, are associated with significant differences in the treatment-induced clearance of the virus. This finding, originally reported in Nature,[31] showed that genotype 1 hepatitis C patients carrying certain genetic variant alleles near the IL28B gene are more possibly to achieve sustained virological response after the treatment than others. Later report from Nature[32] demonstrated that the same genetic variants are also associated with the natural clearance of the genotype 1 hepatitis C virus.
চিকিৎসা
[সম্পাদনা কৰক]When infection attacks the body, anti-infective drugs can suppress the infection. Several broad types of anti-infective drugs exist, depending on the type of organism targeted; they include antibacterial (antibiotic; including antitubercular), antiviral, antifungal and antiparasitic (including antiprotozoal and antihelminthic) agents. Depending on the severity and the type of infection, the antibiotic may be given by mouth or by injection, or may be applied topically. Severe infections of the brain are usually treated with intravenous antibiotics. Sometimes, multiple antibiotics are used in case there is resistance to one antibiotic. Antibiotics only work for bacteria and do not affect viruses. Antibiotics work by slowing down the multiplication of bacteria or killing the bacteria. The most common classes of antibiotics used in medicine include penicillin, cephalosporins, aminoglycosides, macrolides, quinolones and tetracyclines.[উদ্ধৃতিৰ প্ৰয়োজন]
Not all infections require treatment, and for many self-limiting infections the treatment may cause more side-effects than benefits. Antimicrobial stewardship is the concept that healthcare providers should treat an infection with an antimicrobial that specifically works well for the target pathogen for the shortest amount of time and to only treat when there is a known or highly suspected pathogen that will respond to the medication.[33]
ৰোগ বিতৰণ
[সম্পাদনা কৰক]

In 2010 about 10 million people died of an infectious disease.[35]
The World Health Organization collects information on global deaths by International Classification of Disease (ICD) code categories. The following table lists the top infectious disease by number of deaths in 2002. 1993 data is included for comparison.
| Rank | Cause of death | Deaths 2002 (in millions) |
Percentage of all deaths |
Deaths 1993 (in millions) |
1993 Rank |
|---|---|---|---|---|---|
| N/A | All infectious diseases | 14.7 | 25.9% | 16.4 | 32.2% |
| 1 | Lower respiratory infections[38] | 3.9 | 6.9% | 4.1 | 1 |
| 2 | HIV/AIDS | 2.8 | 4.9% | 0.7 | 7 |
| 3 | Diarrheal diseases[39] | 1.8 | 3.2% | 3.0 | 2 |
| 4 | Tuberculosis (TB) | 1.6 | 2.7% | 2.7 | 3 |
| 5 | Malaria | 1.3 | 2.2% | 2.0 | 4 |
| 6 | Measles | 0.6 | 1.1% | 1.1 | 5 |
| 7 | Pertussis | 0.29 | 0.5% | 0.36 | 7 |
| 8 | Tetanus | 0.21 | 0.4% | 0.15 | 12 |
| 9 | Meningitis | 0.17 | 0.3% | 0.25 | 8 |
| 10 | Syphilis | 0.16 | 0.3% | 0.19 | 11 |
| 11 | Hepatitis B | 0.10 | 0.2% | 0.93 | 6 |
| 12-17 | Tropical diseases (6)[40] | 0.13 | 0.2% | 0.53 | 9, 10, 16–18 |
| Note: Other causes of death include maternal and perinatal conditions (5.2%), nutritional deficiencies (0.9%), noncommunicable conditions (58.8%), and injuries (9.1%). | |||||
The top three single agent/disease killers are HIV/AIDS, TB and malaria. While the number of deaths due to nearly every disease have decreased, deaths due to HIV/AIDS have increased fourfold. Childhood diseases include pertussis, poliomyelitis, diphtheria, measles and tetanus. Children also make up a large percentage of lower respiratory and diarrheal deaths. In 2012, approximately 3.1 million people have died due to lower respiratory infections, making it the number 4 leading cause of death in the world.[41]
Historic pandemics
[সম্পাদনা কৰক]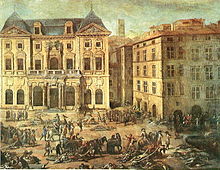
A pandemic (or global epidemic) is a disease that affects people over an extensive geographical area.
- Plague of Justinian, from 541 to 542, killed between 50% and 60% of Europe's population.[42]
- The Black Death of 1347 to 1352 killed 25 million in Europe over 5 years. The plague reduced the old world population from an estimated 450 million to between 350 and 375 million in the 14th century.
- The introduction of smallpox, measles, and typhus to the areas of Central and South America by European explorers during the 15th and 16th centuries caused pandemics among the native inhabitants. Between 1518 and 1568 disease pandemics are said to have caused the population of Mexico to fall from 20 million to 3 million.[43]
- The first European influenza epidemic occurred between 1556 and 1560, with an estimated mortality rate of 20%.[43]
- Smallpox killed an estimated 60 million Europeans during the 18th century[44] (approximately 400,000 per year).[45] Up to 30% of those infected, including 80% of the children under 5 years of age, died from the disease, and one-third of the survivors went blind.[46]
- In the 19th century, tuberculosis killed an estimated one-quarter of the adult population of Europe;[47] by 1918 one in six deaths in France were still caused by TB.
- The Influenza Pandemic of 1918 (or the Spanish Flu) killed 25–50 million people (about 2% of world population of 1.7 billion).[48] Today Influenza kills about 250,000 to 500,000 worldwide each year.
Emerging diseases
[সম্পাদনা কৰক]In most cases, microorganisms live in harmony with their hosts via mutual or commensal interactions. Diseases can emerge when existing parasites become pathogenic or when new pathogenic parasites enter a new host.
- Coevolution between parasite and host can lead to hosts becoming resistant to the parasites or the parasites may evolve greater virulence, leading to immunopathological disease.
- Human activity is involved with many emerging infectious diseases, such as environmental change enabling a parasite to occupy new niches. When that happens, a pathogen that had been confined to a remote habitat has a wider distribution and possibly a new host organism. Parasites jumping from nonhuman to human hosts are known as zoonoses. Under disease invasion, when a parasite invades a new host species, it may become pathogenic in the new host.[49]
Several human activities have led to the emergence of zoonotic human pathogens, including viruses, bacteria, protozoa, and rickettsia,[50] and spread of vector-borne diseases,[49] see also globalization and disease and wildlife disease:
- Encroachment on wildlife habitats. The construction of new villages and housing developments in rural areas force animals to live in dense populations, creating opportunities for microbes to mutate and emerge.[51]
- Changes in agriculture. The introduction of new crops attracts new crop pests and the microbes they carry to farming communities, exposing people to unfamiliar diseases.
- The destruction of rain forests. As countries make use of their rain forests, by building roads through forests and clearing areas for settlement or commercial ventures, people encounter insects and other animals harboring previously unknown microorganisms.
- Uncontrolled urbanization. The rapid growth of cities in many developing countries tends to concentrate large numbers of people into crowded areas with poor sanitation. These conditions foster transmission of contagious diseases.
- Modern transport. Ships and other cargo carriers often harbor unintended "passengers", that can spread diseases to faraway destinations. While with international jet-airplane travel, people infected with a disease can carry it to distant lands, or home to their families, before their first symptoms appear.
ইতিহাস
[সম্পাদনা কৰক]Ideas of contagion became more popular in Europe during the Renaissance, particularly through the writing of the Italian physician Girolamo Fracastoro.[52]
Anton van Leeuwenhoek (1632–1723) advanced the science of microscopy by being the first to observe microorganisms, allowing for easy visualization of bacteria.
In the mid-19th century John Snow and William Budd did important work demonstrating the contagiousness of typhoid and cholera through contaminated water. Both are credited with decreasing epidemics of cholera in their towns by implementing measures to prevent contamination of water.[53]
Louis Pasteur proved beyond doubt that certain diseases are caused by infectious agents, and developed a vaccine for rabies.
Robert Koch, provided the study of infectious diseases with a scientific basis known as Koch's postulates.
Edward Jenner, Jonas Salk and Albert Sabin developed effective vaccines for smallpox and polio, which would later result in the eradication and near-eradication of these diseases, respectively.
Alexander Fleming discovered the world's first antibiotic, Penicillin, which Florey and Chain then developed.
Gerhard Domagk developed sulphonamides, the first broad spectrum synthetic antibacterial drugs.
Medical specialists
[সম্পাদনা কৰক]The medical treatment of infectious diseases falls into the medical field of Infectious Disease and in some cases the study of propagation pertains to the field of Epidemiology. Generally, infections are initially diagnosed by primary care physicians or internal medicine specialists. For example, an "uncomplicated" pneumonia will generally be treated by the internist or the pulmonologist (lung physician). The work of the infectious diseases specialist therefore entails working with both patients and general practitioners, as well as laboratory scientists, immunologists, bacteriologists and other specialists.
An infectious disease team may be alerted when:
- The disease has not been definitively diagnosed after an initial workup
- The patient is immunocompromised (for example, in AIDS or after chemotherapy);
- The infectious agent is of an uncommon nature (e.g. tropical diseases);
- The disease has not responded to first line antibiotics;
- The disease might be dangerous to other patients, and the patient might have to be isolated
Society and culture
[সম্পাদনা কৰক]A number of studies have reported associations between pathogen load in an area and human behavior. Higher pathogen load is associated with decreased size of ethnic and religious groups in an area. This may be due high pathogen load favoring avoidance of other groups, which may reduce pathogen transmission, or a high pathogen load preventing the creation of large settlements and armies that enforce a common culture. Higher pathogen load is also associated with more restricted sexual behavior, which may reduce pathogen transmission. It also associated with higher preferences for health and attractiveness in mates. Higher fertility rates and shorter or less parental care per child is another association that may be a compensation for the higher mortality rate. There is also an association with polygyny which may be due to higher pathogen load, making selecting males with a high genetic resistance increasingly important. Higher pathogen load is also associated with more collectivism and less individualism, which may limit contacts with outside groups and infections. There are alternative explanations for at least some of the associations although some of these explanations may in turn ultimately be due to pathogen load. Thus, polygny may also be due to a lower male:female ratio in these areas but this may ultimately be due to male infants having increased mortality from infectious diseases. Another example is that poor socioeconomic factors may ultimately in part be due to high pathogen load preventing economic development.[54]
Fossil record
[সম্পাদনা কৰক]
Evidence of infection in fossil remains is a subject of interest for paleopathologists, scientists who study occurrences of injuries and illness in extinct life forms. Signs of infection have been discovered in the bones of carnivorous dinosaurs. When present, however, these infections seem to tend to be confined to only small regions of the body. A skull attributed to the early carnivorous dinosaur Herrerasaurus ischigualastensis exhibits pit-like wounds surrounded by swollen and porous bone. The unusual texture of the bone around the wounds suggests they were afflicted by a short-lived, non-lethal infection. Scientists who studied the skull speculated that the bite marks were received in a fight with another Herrerasaurus. Other carnivorous dinosaurs with documented evidence of infection include Acrocanthosaurus, Allosaurus, Tyrannosaurus and a tyrannosaur from the Kirtland Formation. The infections from both tyrannosaurs were received by being bitten during a fight, like the Herrerasaurus specimen.[55]
Outer space
[সম্পাদনা কৰক]A 2006 Space Shuttle experiment found that Salmonella typhimurium, a bacterium that can cause food poisoning, became more virulent when cultivated in space.[56] On April 29, 2013, scientists in Rensselaer Polytechnic Institute, funded by NASA, reported that, during spaceflight on the International Space Station, microbes seem to adapt to the space environment in ways "not observed on Earth" and in ways that "can lead to increases in growth and virulence".[57] More recently, in 2017, bacteria were found to be more resistant to antibiotics and to thrive in the near-weightlessness of space.[58] Microorganisms have been observed to survive the vacuum of outer space.[59][60]
আৰু চাওক
[সম্পাদনা কৰক]- Bioinformatics Resource Centers for Infectious Diseases
- Biological hazard
- Blood-borne disease
- Coinfection
- Copenhagen Consensus
- Cordon sanitaire
- Disease diffusion mapping
- Foodborne illness
- History of medicine
- Hospital-acquired infection
- Eradication of infectious diseases
- Human Microbiome Project
- Infection control
- Isolation (health care)
- List of bacterial vaginosis microbiota
- List of causes of death by rate
- List of diseases caused by insects
- List of epidemics
- List of infectious diseases
- Mathematical modelling of infectious disease
- Membrane vesicle trafficking
- Multiplicity of infection
- Neglected tropical diseases
- Social distancing
- Spatiotemporal Epidemiological Modeler (STEM)
- Spillover infection
- Threshold host density
- Transmission (medicine)
- Ubi pus, ibi evacua (Latin: "where there is pus, there evacuate it")
- Vaccine-preventable diseases
- Waterborne diseases
-->
টোকা আৰু তথ্য উৎস
[সম্পাদনা কৰক]- ↑ Definition of "infection" from several medical dictionaries – Retrieved on 2012-04-03
- ↑ "Utilizing antibiotics agents effectively will preserve present day medication". News Ghana. 21 November 2015. http://newsghana.com.gh/?p=853675। আহৰণ কৰা হৈছে: 21 November 2015.
- ↑ Alberto Signore (2013). "About inflammation and infection". EJNMMI Research খণ্ড 8 (3). http://www.ejnmmires.com/content/pdf/2191-219X-3-8.pdf.
- ↑ GBD 2013 Mortality and Causes of Death, Collaborators (17 December 2014). "Global, regional, and national age-sex specific all-cause and cause-specific mortality for 240 causes of death, 1990-2013: a systematic analysis for the Global Burden of Disease Study 2013.". Lancet খণ্ড 385 (9963): 117–71. doi:10.1016/S0140-6736(14)61682-2. PMID 25530442.
- ↑ "Infectious Disease, Internal Medicine". Association of American Medical Colleges. Archived from the original on 2015-02-06. https://web.archive.org/web/20150206201010/https://www.aamc.org/cim/specialty/list/us/339608/infectious_disease_-internal_medicine.html। আহৰণ কৰা হৈছে: 2015-08-20. "Infectious disease is the subspecialty of internal medicine dealing with the diagnosis and treatment of communicable diseases of all types, in all organs, and in all ages of patients."
- ↑ 6.0 6.1 This section incorporates public domain materials included in the text: Medical Microbiology Fourth Edition: Chapter 8 (1996). Baron, Samuel MD. The University of Texas Medical Branch at Galveston. "Archived copy". Archived from the original on June 29, 2009. https://web.archive.org/web/20090629132331/http://www.ncbi.nlm.nih.gov/books/bv.fcgi?rid=mmed। আহৰণ কৰা হৈছে: 2013-11-27.
- ↑ 7.0 7.1 7.2 7.3 7.4 7.5 Ryan KJ; Ray CG (editors) (2004). Sherris Medical Microbiology (4th সম্পাদনা). McGraw Hill. ISBN 0-8385-8529-9.
- ↑ (Higurea & Pietrangelo 2016).
- ↑ 9.0 9.1 9.2 "Bacterial vs. Viral Infections – Do You Know the Difference?" National Information Program on Antibiotics
- ↑ Robert N. Golden; Fred Peterson. The Truth About Illness and Disease. Infobase Publishing, 2009. পৃষ্ঠা. 181. ISBN 1438126379.
- ↑ "Infection". Rencare. Archived from the original on March 5, 2012. https://web.archive.org/web/20120305023556/http://www.rencareltd.com/conditions/infections/। আহৰণ কৰা হৈছে: 4 July 2013.
- ↑ Infection Cycle – Retrieved on 2010-01-21 Archived May 17, 2014, at the Wayback Machine
- ↑ Understanding Infectious Diseases Archived 2009-09-24 at the Wayback Machine Science.Education.Nih.Gov article – Retrieved on 2010-01-21
- ↑ Duerkop, Breck A; Hooper, Lora V (2013-07-01). "Resident viruses and their interactions with the immune system" (en ভাষাত). Nature Immunology খণ্ড 14 (7): 654–59. doi:10.1038/ni.2614. PMID 23778792. PMC 3760236. http://www.nature.com/ni/journal/v14/n7/fig_tab/ni.2614_F2.html.
- ↑ "Bacterial Pathogenesis at Washington University". প্ৰকাশক St. Louis: StudyBlue. https://www.studyblue.com/notes/note/n/bacterial-pathogenesis/deck/11094651। আহৰণ কৰা হৈছে: 2016-12-02.
- ↑ "Print Friendly". www.lifeextension.com. http://www.lifeextension.com/magazine/2014/6/the-dangers-of-using-antibiotics-to-prevent-urinary-tract-infections/page-01?p=1। আহৰণ কৰা হৈছে: 2016-12-02.
- ↑ "Does this patient have an infection of a chronic wound?". JAMA খণ্ড 307 (6): 605–11. Feb 2012. doi:10.1001/jama.2012.98. PMID 22318282.
- ↑ http://www.immunize.org/catg.d/p4215.pdf
- ↑ Chronic Infection Information Retrieved on 2010-01-14 Archived July 22, 2015, at the Wayback Machine
- ↑ Intestinal Parasites and Infection Archived 2010-10-28 at the Wayback Machine fungusfocus.com – Retrieved on 2010-01-21
- ↑ "Clinical Infectious Disease – Introduction". www.microbiologybook.org. http://www.microbiologybook.org/Infectious%20Disease/Infectious%20Disease%20Introduction.htm। আহৰণ কৰা হৈছে: 2017-04-19.
- ↑ Pathogens and vectors Archived 2017-10-05 at the Wayback Machine. MetaPathogen.com.
- ↑ Ljubin-Sternak, Suncanica; Mestrovic, Tomislav (2014). "Review: Clamydia trachonmatis and Genital Mycoplasmias: Pathogens with an Impact on Human Reproductive Health". Journal of Pathogens খণ্ড 2014 (183167): 3. doi:10.1155/2014/183167. PMID 25614838.
- ↑ 24.0 24.1 "Diagnostic value of clinical features at presentation to identify serious infection in children in developed countries: a systematic review". Lancet খণ্ড 375 (9717): 834–45. March 2010. doi:10.1016/S0140-6736(09)62000-6. PMID 20132979.
- ↑ "The effectiveness of hand hygiene procedures including hand-washing and alcohol-based hand sanitizers in reducing the risks of infections in home and community settings". American Journal of Infection Control খণ্ড 35 (10): S27–S64. 2007. doi:10.1016/j.ajic.2007.07.001. http://www.ajicjournal.org/article/S0196-6553(07)00595-0/fulltext.
- ↑ ""Generalized Infectious Cycle" Diagram Illustration". science.education.nih.gov. Archived from the original on 2009-09-24. https://web.archive.org/web/20090924155114/https://science.education.nih.gov/supplements/nih1/diseases/guide/fig4-longdesc.htm। আহৰণ কৰা হৈছে: 2010-01-21.
- ↑ 27.0 27.1 Watts, Duncan (2003). Six degrees: the science of a connected age. প্ৰকাশক London: William Heinemann. ISBN 0-393-04142-5.
- ↑ Preston, Richard (1995). The hot zone. প্ৰকাশক Garden City, N.Y.: Anchor Books. ISBN 0-385-49522-6.
- ↑ "The foot-and-mouth epidemic in Great Britain: pattern of spread and impact of interventions". Science খণ্ড 292 (5519): 1155–60. May 2001. doi:10.1126/science.1061020. PMID 11303090. http://www.sciencemag.org/cgi/pmidlookup?view=long&pmid=11303090.
- ↑ "Infectious diseases epidemiology". Journal of Epidemiology and Community Health খণ্ড 60 (3): 192–95. 2006. doi:10.1136/jech.2003.011593. PMID 16476746.
- ↑ "Genetic variation in IL28B predicts hepatitis C treatment-induced viral clearance". Nature খণ্ড 461 (7262): 399–401. 2009. doi:10.1038/nature08309. PMID 19684573.
- ↑ "Genetic variation in IL28B and spontaneous clearance of hepatitis C virus". Nature খণ্ড 461 (7265): 798–801. 2009. doi:10.1038/nature08463. PMID 19759533.
- ↑ O’Brien, Deirdre J.; Gould, Ian M. (August 2013). "Maximizing the impact of antimicrobial stewardship". Current Opinion in Infectious Diseases খণ্ড 26 (4): 352–58. doi:10.1097/QCO.0b013e3283631046. PMID 23806898.
- ↑ World Health Organization (February 2009). "Age-standardized DALYs per 100,000 by cause, and Member State, 2004". http://www.who.int/entity/healthinfo/global_burden_disease/gbddeathdalycountryestimates2004.xls.
- ↑ "Could Ebola rank among the deadliest communicable diseases?". CBC News. 20 October 2014. http://www.cbc.ca/news/1.2802071.
- ↑ "The World Health Report (Annex Table 2)" (PDF). 2004. http://www.who.int/whr/2004/annex/topic/en/annex_2_en.pdf.
- ↑ "Table 5" (PDF). 1995. http://www.who.int/whr/1995/en/whr95_ch1_en.pdf.
- ↑ Lower respiratory infections include various pneumonias, influenzas and acute bronchitis.
- ↑ Diarrheal diseases are caused by many different organisms, including cholera, botulism, and E. coli to name a few. See also: Intestinal infectious diseases
- ↑ Tropical diseases include Chagas disease, dengue fever, lymphatic filariasis, leishmaniasis, onchocerciasis, schistosomiasis and trypanosomiasis.
- ↑ "WHO | The top 10 causes of death". WHO. http://www.who.int/mediacentre/factsheets/fs310/en/। আহৰণ কৰা হৈছে: 2015-09-24.
- ↑ "Infectious and Epidemic Disease in History" Archived July 12, 2012, at Archive.today
- ↑ 43.0 43.1 "Infectious Diseases and Human Population History". BioScience খণ্ড 46 (2): 115–26. 1996. doi:10.2307/1312814. Archived from the original on 2016-03-03. https://web.archive.org/web/20160303231520/http://www.erin.utoronto.ca/~w3gwynne/BIO418/Dobson1996.pdf। আহৰণ কৰা হৈছে: 2018-02-28.
- ↑ "Smallpox" Archived 2018-01-29 at the Wayback Machine. North Carolina Digital History.
- ↑ Smallpox and Vaccinia. National Center for Biotechnology Information. Archived June 1, 2009, at the Wayback Machine
- ↑ "Smallpox: The Triumph over the Most Terrible of the Ministers of Death"
- ↑ Multidrug-Resistant "Tuberculosis". Centers for Disease Control and Prevention. Archived March 9, 2010, at the Wayback Machine
- ↑ "Influenza of 1918 (Spanish Flu) and the US Navy"
- ↑ 49.0 49.1 Krauss H; Weber A; Appel M (2003). Zoonoses: Infectious Diseases Transmissible from Animals to Humans (3rd সম্পাদনা). প্ৰকাশক Washington, D.C.: ASM Press. ISBN 1-55581-236-8.
- ↑ Potter P (July 2013). "Summer buzz". Emerg Infect Dis খণ্ড 19 (3): 1184. doi:10.3201/eid1907.AC1907. PMC 3903457. http://wwwnc.cdc.gov/eid/article/19/7/ac-1907_article.htm?s_cid=eid_cover_img#suggestedcitation.
- ↑ Peter Daszak; Andrew A. Cunningham; Alex D. Hyatt (27 January 2000). "Emerging Infectious Diseases of Wildlife—Threats to Biodiversity and Human Health". Science খণ্ড 287 (5452): 443–49. doi:10.1126/science.287.5452.443. PMID 10642539. http://science.sciencemag.org/content/287/5452/443.full.
- ↑ Beretta M (2003). "The revival of Lucretian atomism and contagious diseases during the renaissance". Medicina nei secoli খণ্ড 15 (2): 129–54. PMID 15309812.
- ↑ Moorhead Robert (November 2002). "William Budd and typhoid fever". J R Soc Med খণ্ড 95 (11): 561–64. doi:10.1258/jrsm.95.11.561. PMID 12411628.
- ↑ Nettle D (2009). "Ecological influences on human behavioural diversity: a review of recent findings". Trends Ecol. Evol. খণ্ড 24 (11): 618–24. doi:10.1016/j.tree.2009.05.013. PMID 19683831.
- ↑ Molnar, R. E., 2001, "Theropod paleopathology: a literature survey": In: Mesozoic Vertebrate Life, edited by Tanke, D. H., and Carpenter, K., Indiana University Press, pp. 337–63.
- ↑ Caspermeyer, Joe (23 September 2007). "Space flight shown to alter ability of bacteria to cause disease". Arizona State University. Archived from the original on 14 September 2017. https://web.archive.org/web/20170914172718/https://biodesign.asu.edu/news/space-flight-shown-alter-ability-bacteria-cause-disease। আহৰণ কৰা হৈছে: 14 September 2017.
- ↑ "Spaceflight Promotes Biofilm Formation by Pseudomonas aeruginosa". PLOS One খণ্ড 8 (4): e6237. April 29, 2013. doi:10.1371/journal.pone.0062437. http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0062437। আহৰণ কৰা হৈছে: July 5, 2013.
- ↑ Dvorsky, George (13 September 2017). "Alarming Study Indicates Why Certain Bacteria Are More Resistant to Drugs in Space". Gizmodo. https://gizmodo.com/alarming-study-indicates-why-certain-bacteria-are-more-1805666249। আহৰণ কৰা হৈছে: 14 September 2017.
- ↑ Dose, K.; Bieger-Dose, A.; Dillmann, R.; Gill, M.; Kerz, O.; Klein, A.; Meinert, H.; Nawroth, T. et al. (1995). "ERA-experiment "space biochemistry"". Advances in Space Research খণ্ড 16 (8): 119–29. doi:10.1016/0273-1177(95)00280-R. PMID 11542696.
- ↑ Vaisberg, Horneck G.; Eschweiler, U.; Reitz, G.; Wehner, J.; Willimek, R.; Strauch, K. (1995). "Biological responses to space: results of the experiment "Exobiological Unit" of ERA on EURECA I". Adv Space Res. খণ্ড 16 (8): 105–18. doi:10.1016/0273-1177(95)00279-N. PMID 11542695.
বাহ্যিক সংযোগ
[সম্পাদনা কৰক]| বৰ্গীকৰণ |
|---|
| ৱিকিমিডিয়া কমন্সত Infectious diseases and disorders সম্পৰ্কীয় মিডিয়া ফাইল আছে। |
- European Center for Disease Prevention and Control
- U.S. Centers for Disease Control and Prevention,
- Infectious Disease Society of America (IDSA)
- Infectious Disease Index of the Public Health Agency of Canada (PHAC)
- Vaccine Research Center Information concerning vaccine research clinical trials for Emerging and re-Emerging Infectious Diseases.
- Infection Information Resource
- Microbes & Infection (journal)
- Knowledge source for Health Care Professionals involved in Wound management[সংযোগবিহীন উৎস] www.woundsite.info
- Table: Global deaths from communicable diseases, 2010 – Canadian Broadcasting Corp.
